
A Unique and Physiologically Relevant Model
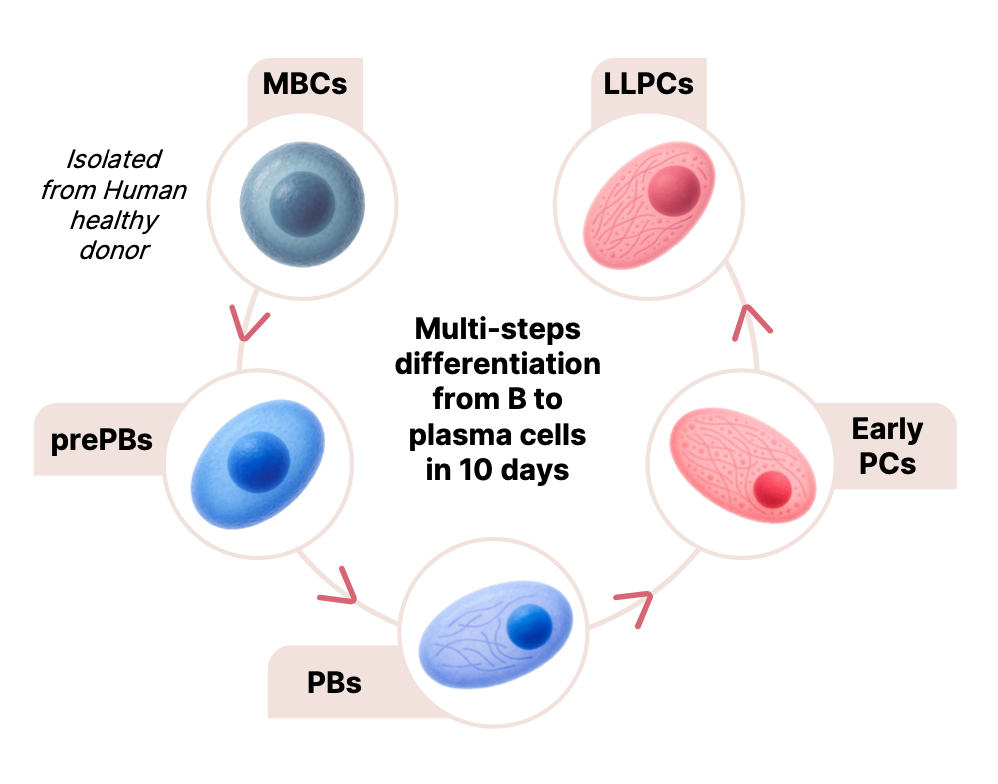
Our multi-step culture system reproduces the sequential differentiation of memory B cells into pre-plasmablasts, plasmablasts, early plasma cells, and fully mature plasma cells using defined combinations of activation signals and cytokines. This approach mimics the differentiation processes occurring across different tissues in vivo. The resulting plasma cells are non-cycling, can survive for several months in vitro, and secrete high levels of immunoglobulins, closely reflecting their in vivo counterparts. These cells express key markers such as CD138 and display gene expression and epigenetic profiles consistent with physiological plasma cells. Transcriptomic and miRNA analyses have further identified key regulatory networks involved in plasma cell differentiation.
Applications in Disease Modeling and Drug Evaluation
This model provides a powerful platform to investigate plasma cells biology and to study therapeutic effects on normal plasma cells generation. It is particularly relevant for understanding disease mechanisms and treatment responses in multiple myeloma and antibody-mediated autoimmune disorders.
Our platform enables the evaluation of drug impact across differentiation stages, including effects on cell survival, differentiation, and immunoglobulin production. It also supports mechanistic studies and biomarker identification to guide therapeutic development.
Deliverables
High-quality, decision-ready data packages including comprehensive reports, as well as raw and processed datasets, supporting preclinical and translational research.